# Load packages.
library(tidyverse)── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
✔ dplyr 1.1.4 ✔ readr 2.1.6
✔ forcats 1.0.1 ✔ stringr 1.6.0
✔ ggplot2 4.0.1 ✔ tibble 3.3.1
✔ lubridate 1.9.4 ✔ tidyr 1.3.2
✔ purrr 1.2.1
── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()
ℹ Use the conflicted package (<http://conflicted.r-lib.org/>) to force all conflicts to become errorslibrary(tidymodels)── Attaching packages ────────────────────────────────────── tidymodels 1.4.1 ──
✔ broom 1.0.12 ✔ rsample 1.3.2
✔ dials 1.4.2 ✔ tailor 0.1.0
✔ infer 1.1.0 ✔ tune 2.0.1
✔ modeldata 1.5.1 ✔ workflows 1.3.0
✔ parsnip 1.4.1 ✔ workflowsets 1.1.1
✔ recipes 1.3.1 ✔ yardstick 1.3.2
── Conflicts ───────────────────────────────────────── tidymodels_conflicts() ──
✖ scales::discard() masks purrr::discard()
✖ dplyr::filter() masks stats::filter()
✖ recipes::fixed() masks stringr::fixed()
✖ dplyr::lag() masks stats::lag()
✖ yardstick::spec() masks readr::spec()
✖ recipes::step() masks stats::step()library(dplyr)
library(ggplot2)
library(here)here() starts at C:/Users/aniss/Desktop/anissawallerdelvalle-portfolio# Load the data and inspect.
data <- read_csv(here("fitting-exercise","Mavoglurant_A2121_nmpk.csv"))Rows: 2678 Columns: 17
── Column specification ────────────────────────────────────────────────────────
Delimiter: ","
dbl (17): ID, CMT, EVID, EVI2, MDV, DV, LNDV, AMT, TIME, DOSE, OCC, RATE, AG...
ℹ Use `spec()` to retrieve the full column specification for this data.
ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.glimpse(data)Rows: 2,678
Columns: 17
$ ID <dbl> 793, 793, 793, 793, 793, 793, 793, 793, 793, 793, 793, 793, 793, …
$ CMT <dbl> 1, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 1, 2, 2, 2, 2, 2,…
$ EVID <dbl> 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0,…
$ EVI2 <dbl> 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0,…
$ MDV <dbl> 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0,…
$ DV <dbl> 0.00, 491.00, 605.00, 556.00, 310.00, 237.00, 147.00, 101.00, 72.…
$ LNDV <dbl> 0.000, 6.196, 6.405, 6.321, 5.737, 5.468, 4.990, 4.615, 4.282, 3.…
$ AMT <dbl> 25, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 25, 0, 0, 0, 0, …
$ TIME <dbl> 0.000, 0.200, 0.250, 0.367, 0.533, 0.700, 1.200, 2.200, 3.200, 4.…
$ DOSE <dbl> 25, 25, 25, 25, 25, 25, 25, 25, 25, 25, 25, 25, 25, 25, 25, 25, 2…
$ OCC <dbl> 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,…
$ RATE <dbl> 75, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 150, 0, 0, 0, 0,…
$ AGE <dbl> 42, 42, 42, 42, 42, 42, 42, 42, 42, 42, 42, 42, 42, 42, 42, 42, 2…
$ SEX <dbl> 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,…
$ RACE <dbl> 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2,…
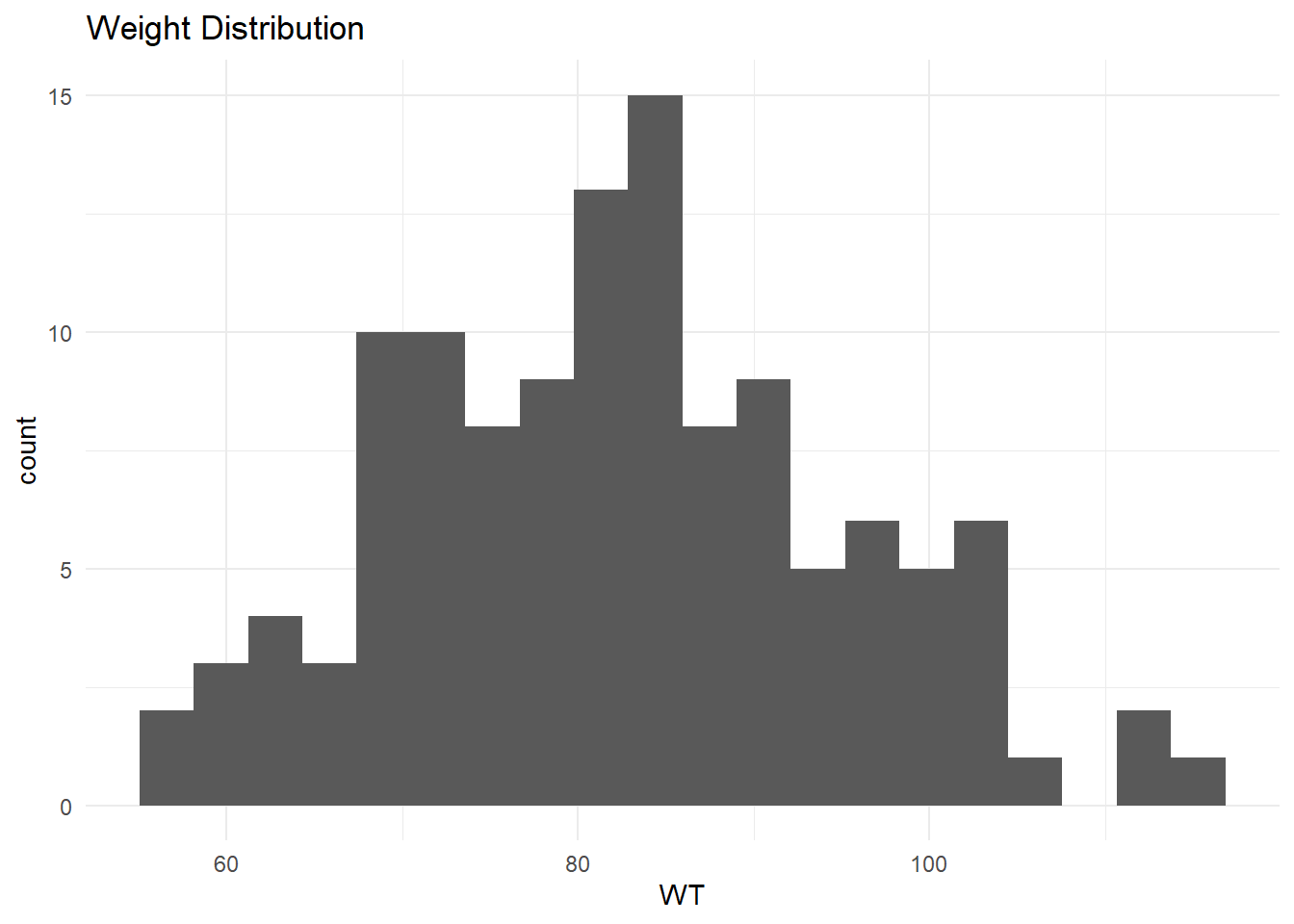
$ WT <dbl> 94.3, 94.3, 94.3, 94.3, 94.3, 94.3, 94.3, 94.3, 94.3, 94.3, 94.3,…
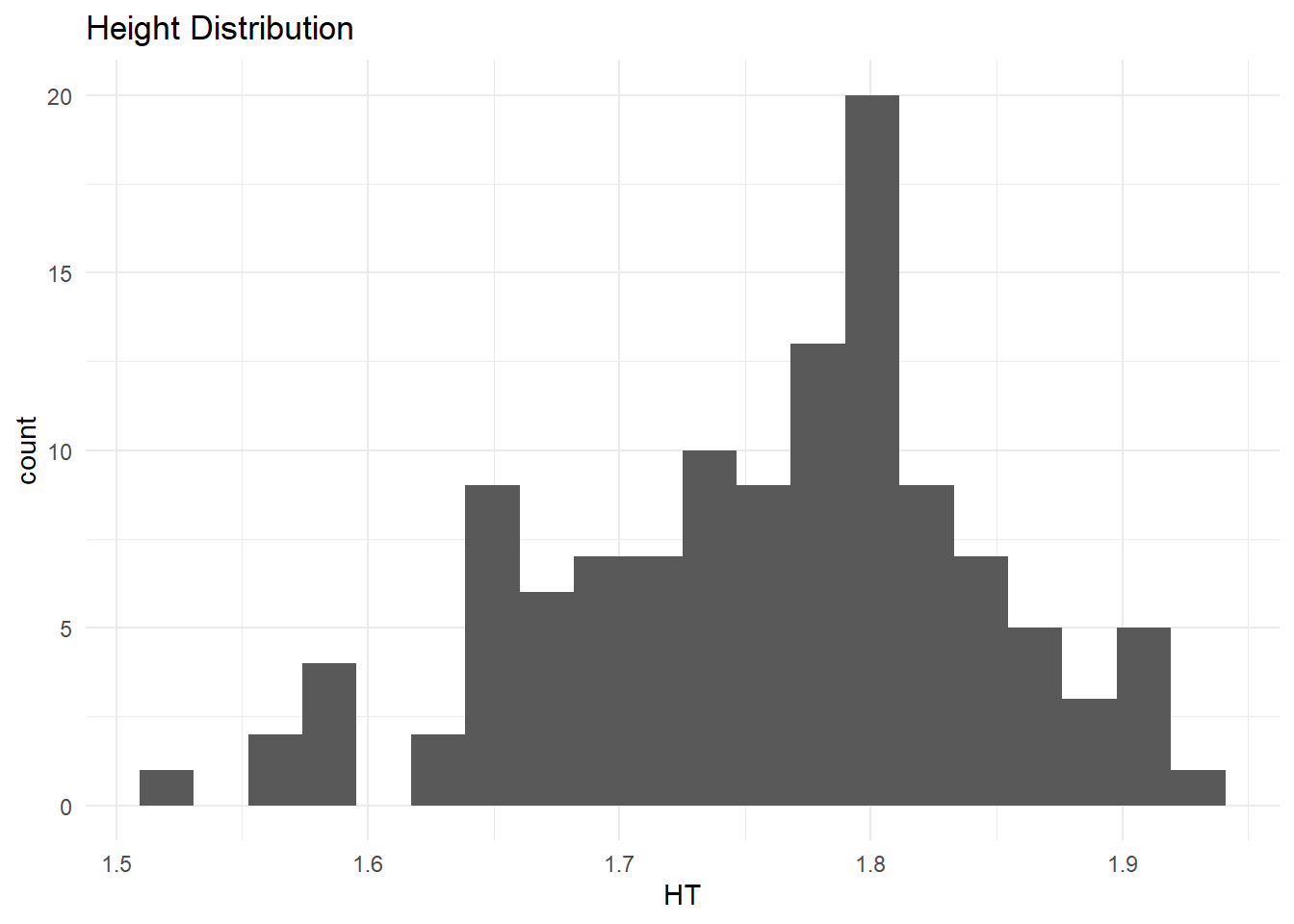
$ HT <dbl> 1.769997, 1.769997, 1.769997, 1.769997, 1.769997, 1.769997, 1.769…