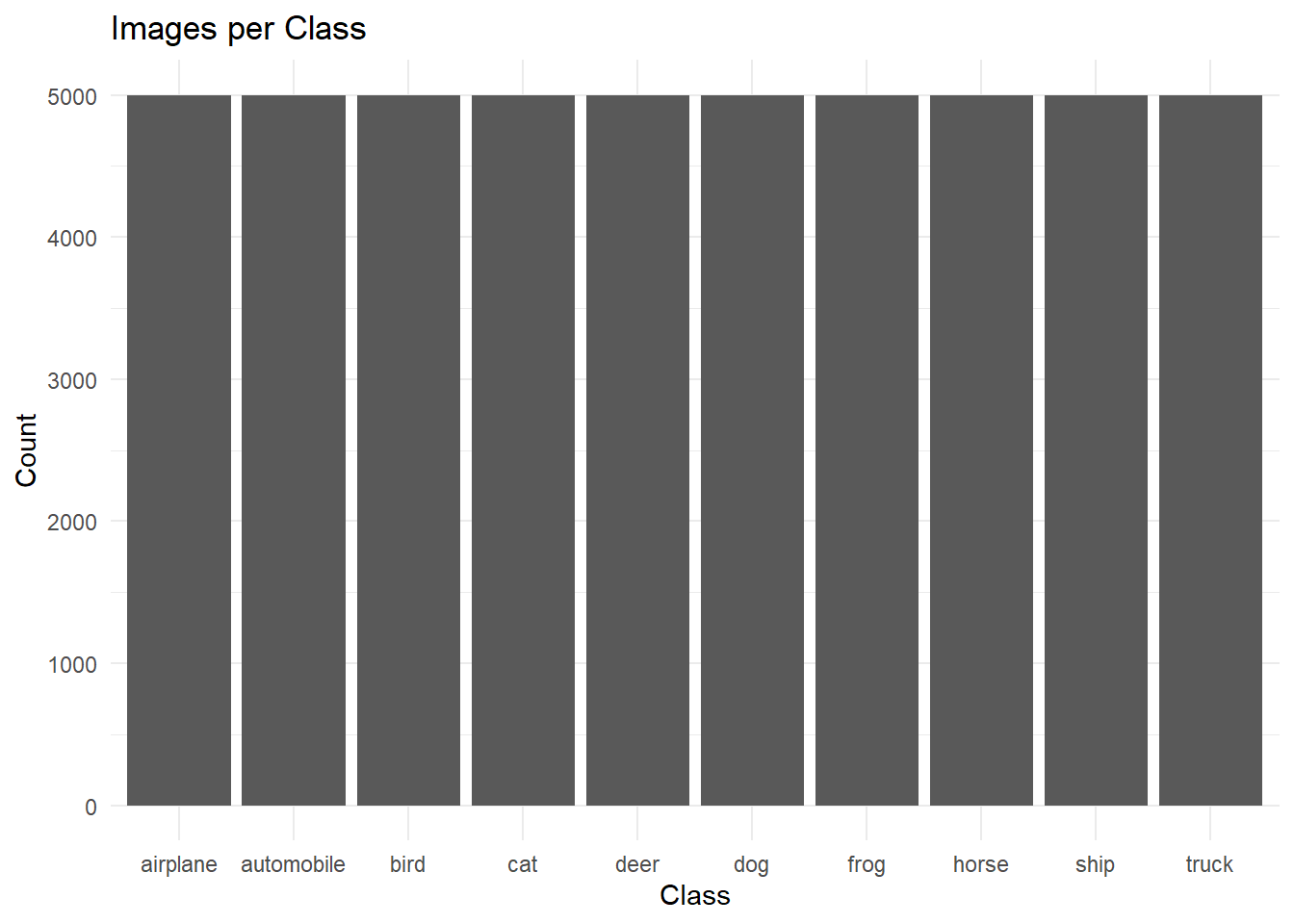# This assignment asks that students analyze a complex dataset. To this end, I will be analyzing the CIFAR-10 image dataset, which consists of 60,000 color images.
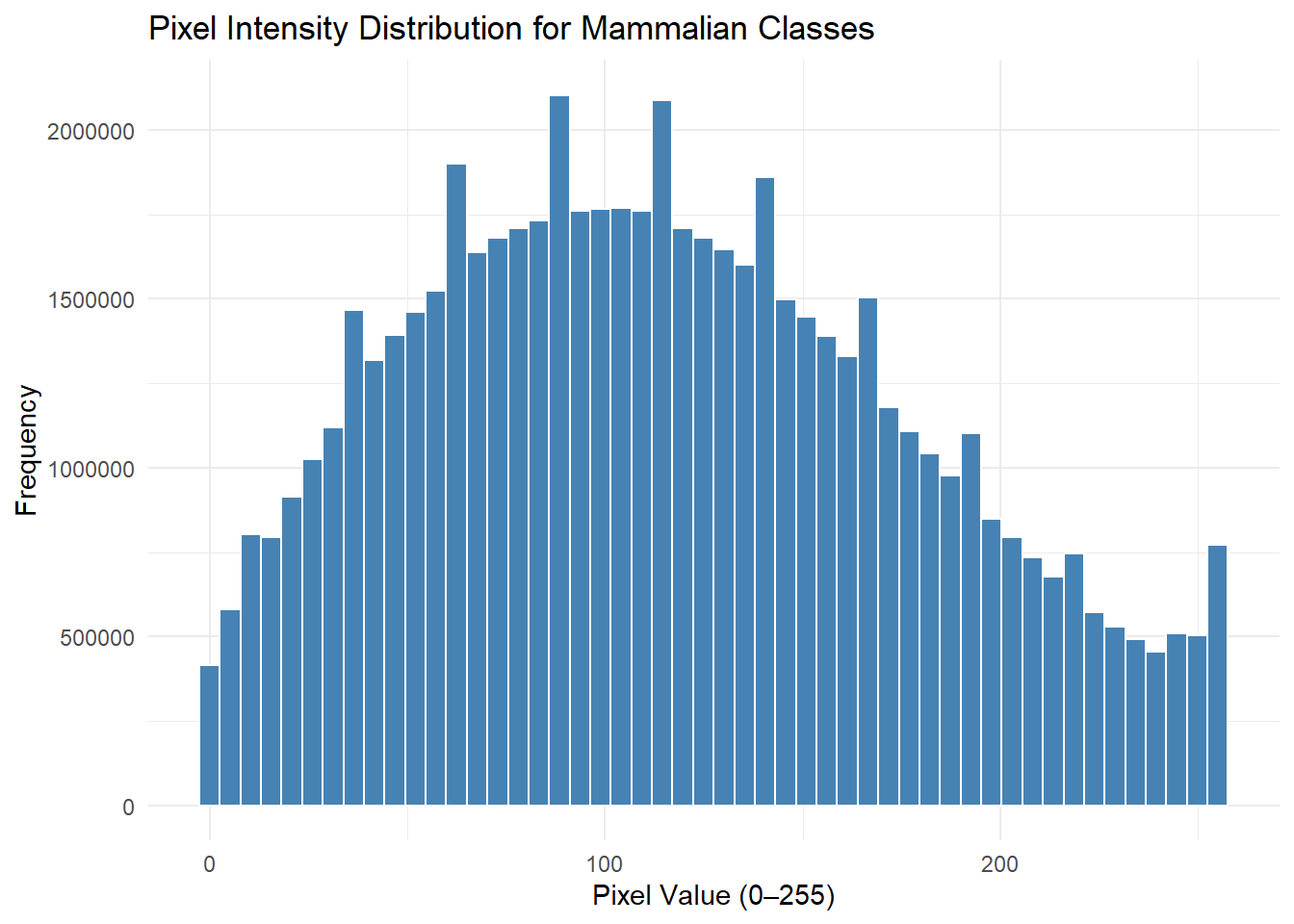
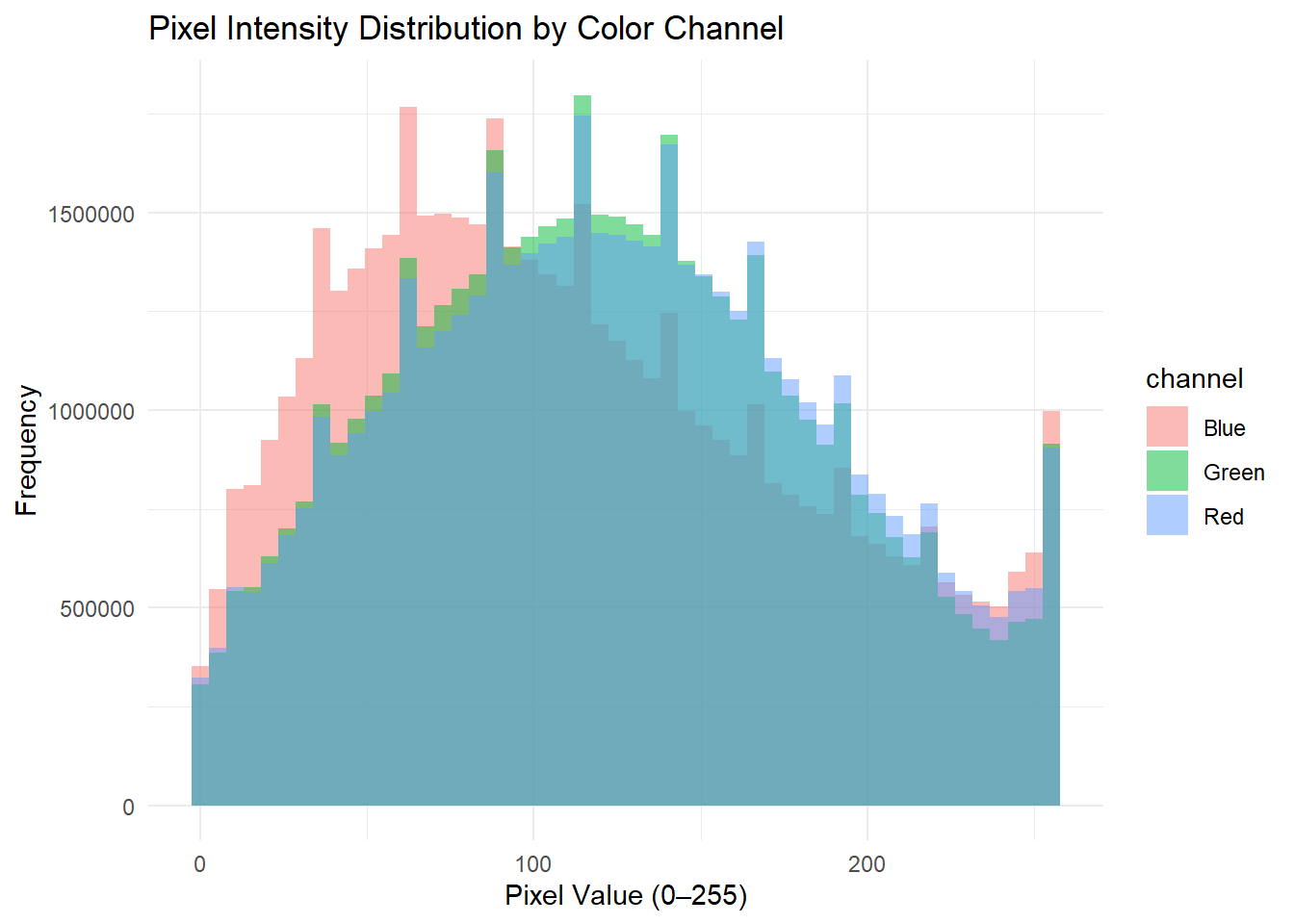
# The CTEGD Cytometry Shared Resource Laboratory at the University of Georgia has a number of instruments -- one such instrument being the Cytek Amnis Imagestream Mk II Imaging Flow Cytometer. I have used this instrument in the past and needed to explore pixel intensity distributions to analyze the data I obtained; this was my motivation for analyzing this dataset for this assignment.
# Install packages. Script below.
# install.packages("keras3", "dplyr", "ggplot2")
# Load libraries.
library("tidyverse")── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
✔ dplyr 1.1.4 ✔ readr 2.1.6
✔ forcats 1.0.1 ✔ stringr 1.6.0
✔ ggplot2 4.0.1 ✔ tibble 3.3.1
✔ lubridate 1.9.4 ✔ tidyr 1.3.2
✔ purrr 1.2.1
── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()
ℹ Use the conflicted package (<http://conflicted.r-lib.org/>) to force all conflicts to become errorslibrary("keras3")
library("dplyr")
library("ggplot2")